The Binomial Generalized Linear Model
Recap of normal linear models
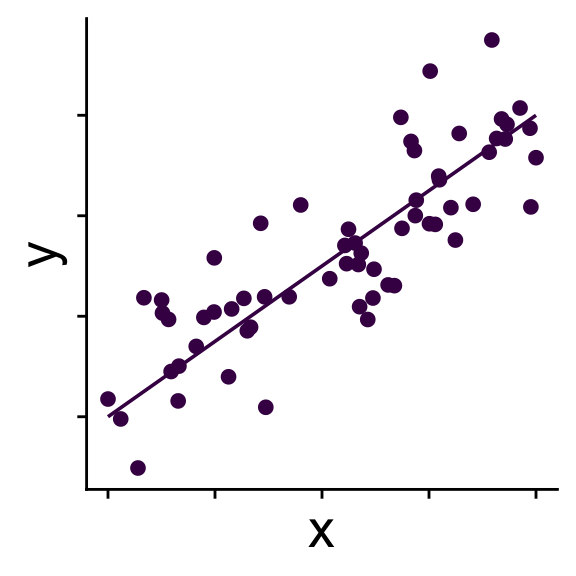
Simple linear model
No random effects
Linear model with random intercepts
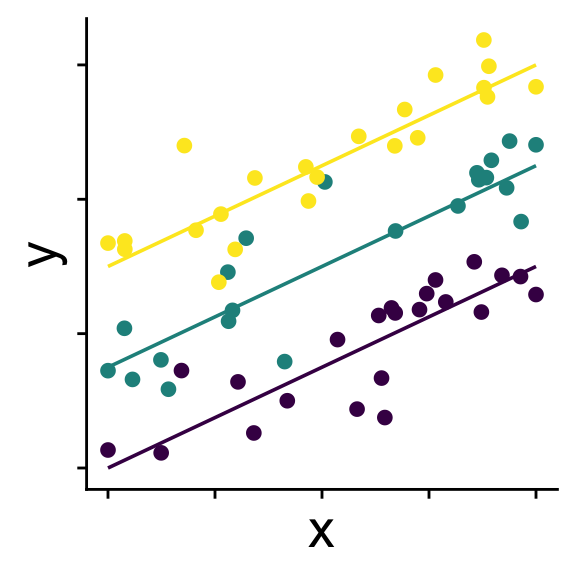
Groups (usually > 3) affect intercepts
Linear model with random slopes
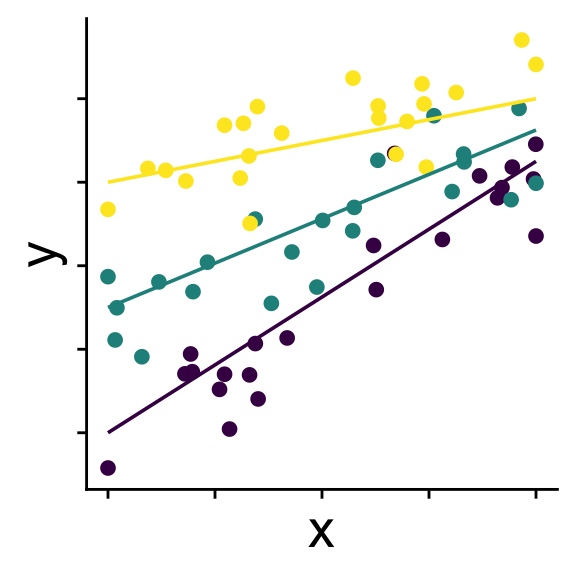
Groups (usually > 3) affect intercepts and slopes
Recap of normal linear models
Simple linear model
\[ \begin{aligned} \bar{y} &= b_0 + b_1 x \\ y_i &\sim \mathcal{N}(\bar{y},~\sigma) \end{aligned} \]
Recap of normal linear models
Linear model with random intercepts
\[ \begin{aligned} r_{0}^{(group~k)} &\sim \mathcal{N}(0,~\sigma_{r_0}) \\ \bar{y} &= b_0 + r_0^{(group~k)} + b_1 x \\ y &\sim \mathcal{N}(\bar{y},~\sigma) \end{aligned} \]
Recap of normal linear models
Linear model with random slopes
\[ \begin{aligned} r_0^{(group~k)} &\sim \mathcal{N}(0,~\sigma_{r_0}) \\ r_1^{(group~k)} &\sim \mathcal{N}(0,~\sigma_{r_1}) \\ \bar{y} &= b_0 + r_0^{(group~k)} + (b_1 + r_1^{(group~k)}) x \\ y &\sim \mathcal{N}(\bar{y},~\sigma) \end{aligned} \]
(technically we usually model a correlation term between \(r_0\) and \(r_1\))
Recap of normal linear models
But what’s so special about normal distribution?
…nothing, we can use other distributions
Introducing: generalized linear models
When we use a distribution other than a normal we call our model a generalized linear model
We’ll start with the binomial as an example
Binomial generalized linear models
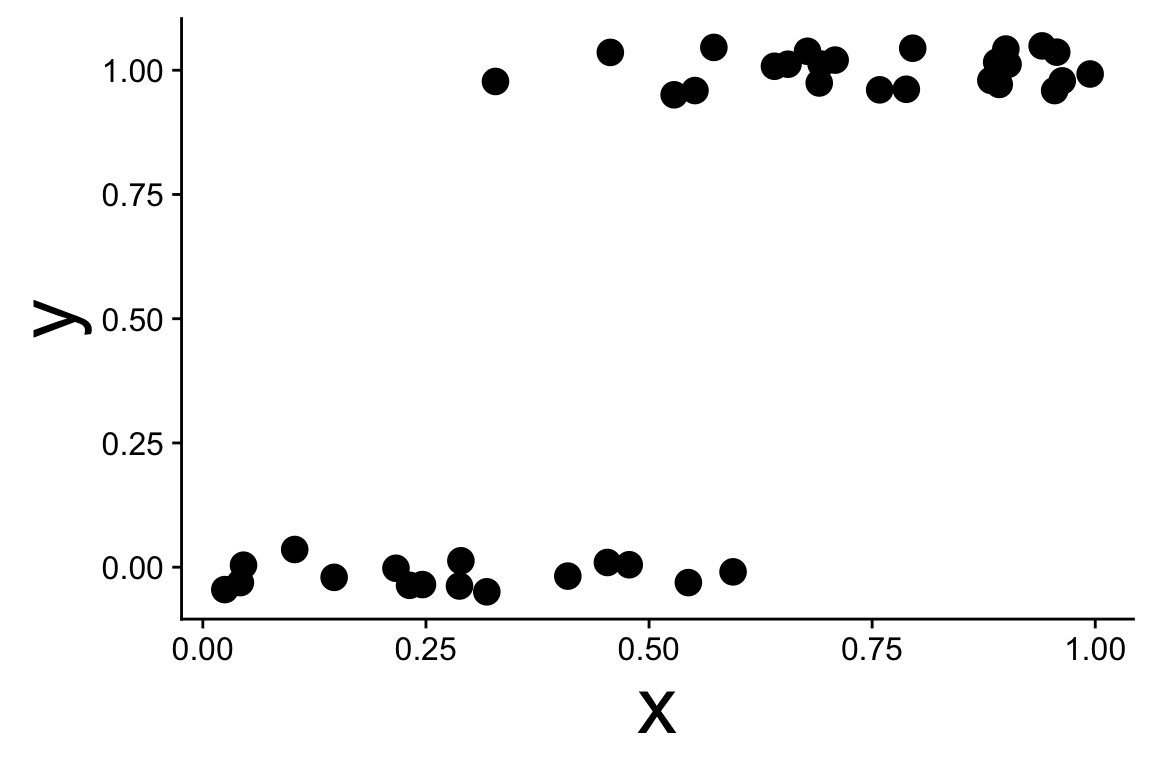
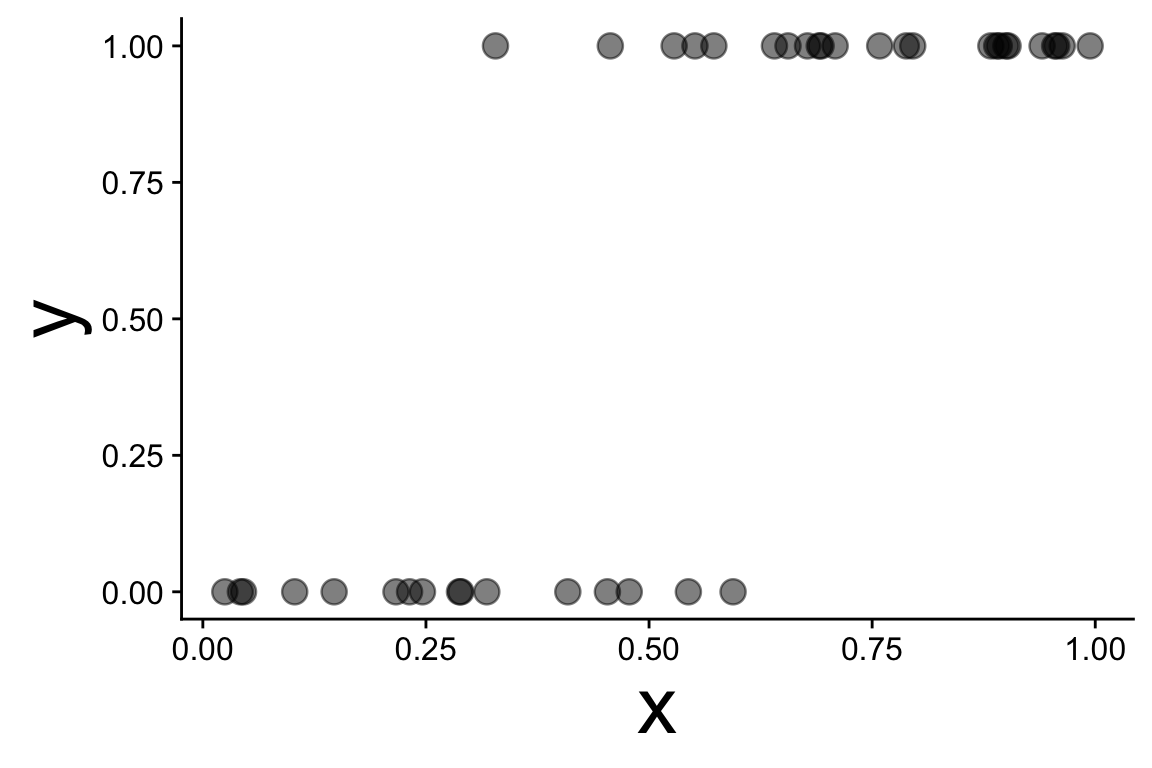
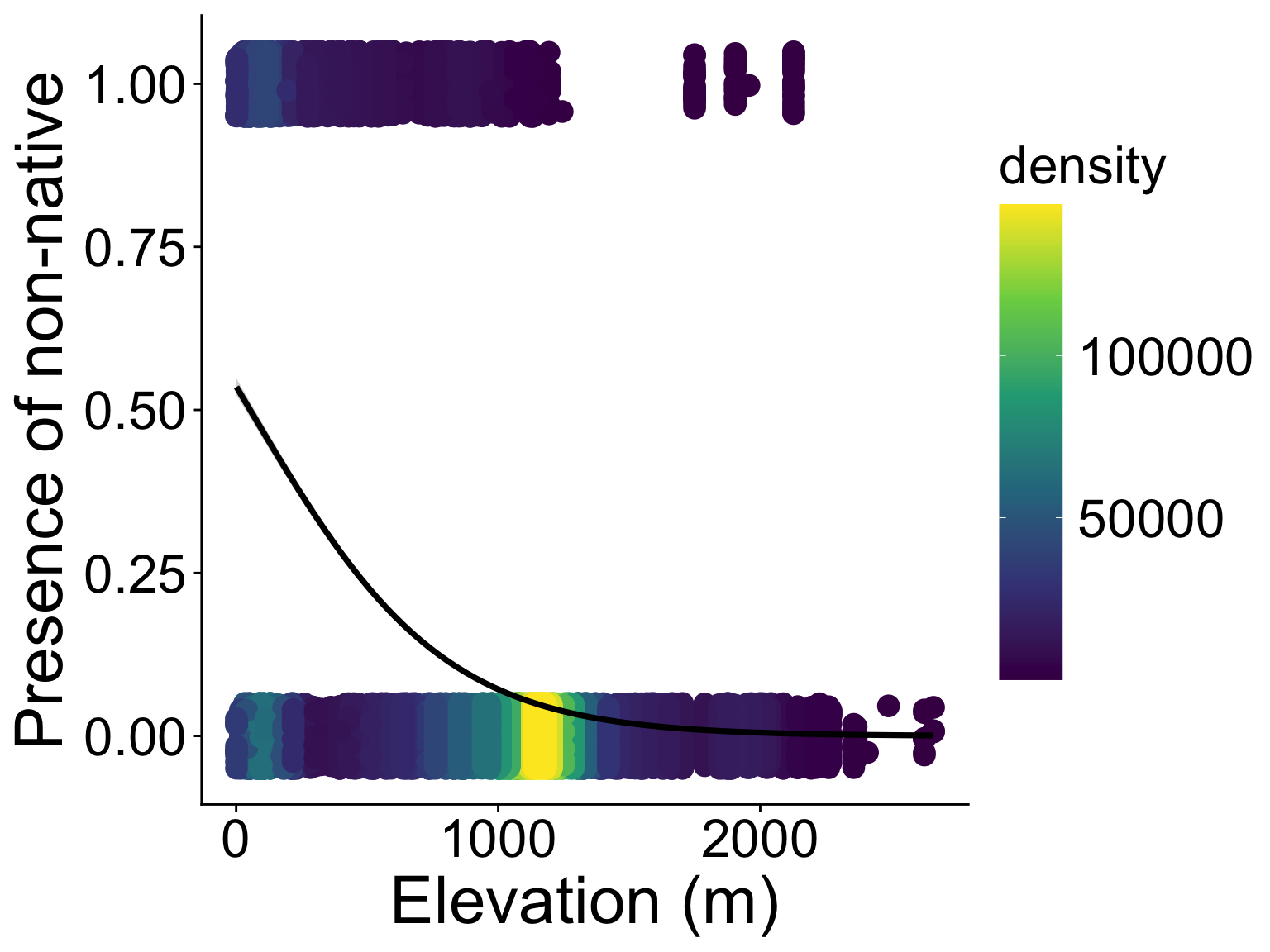
These data are not normal, and fitting a straight line doesn’t make sense
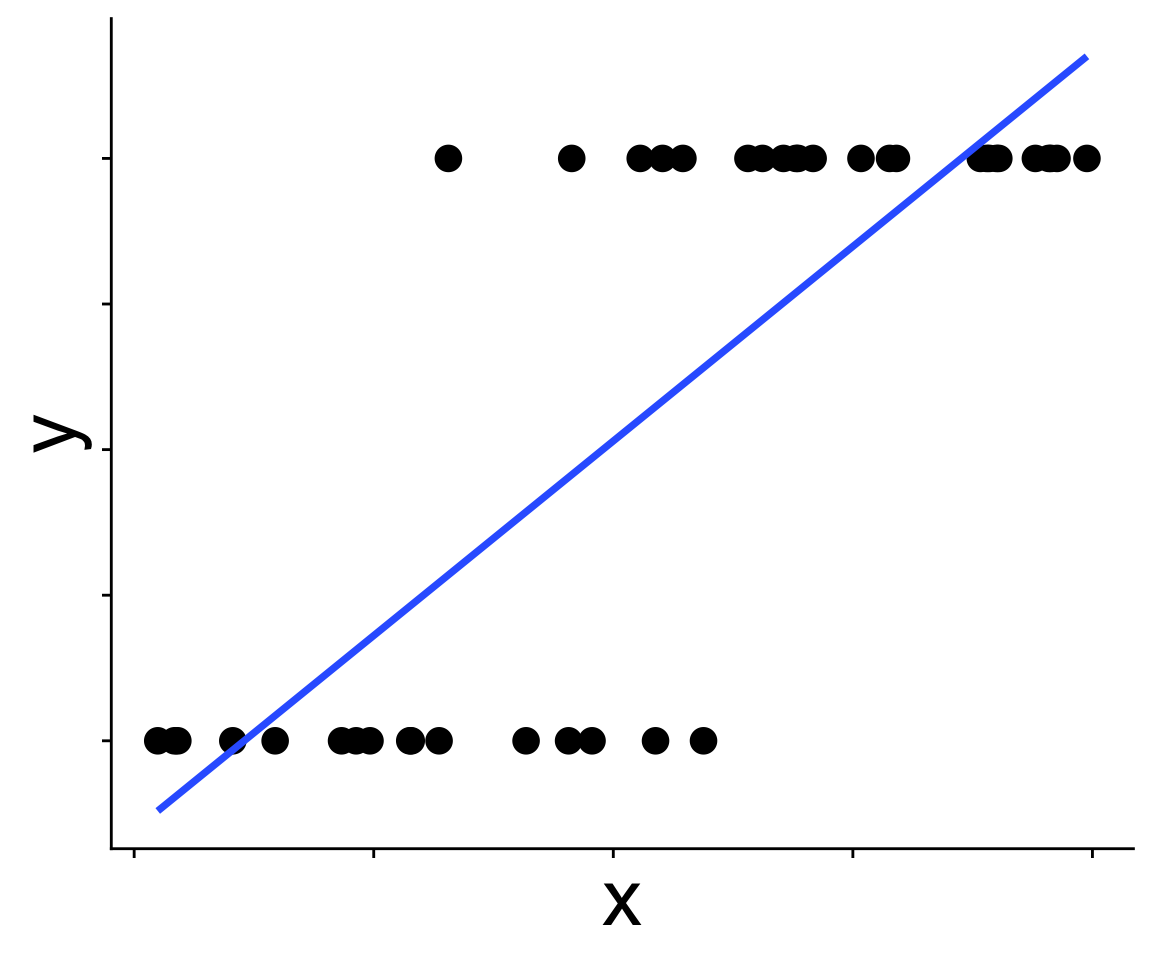
Binomial generalized linear models
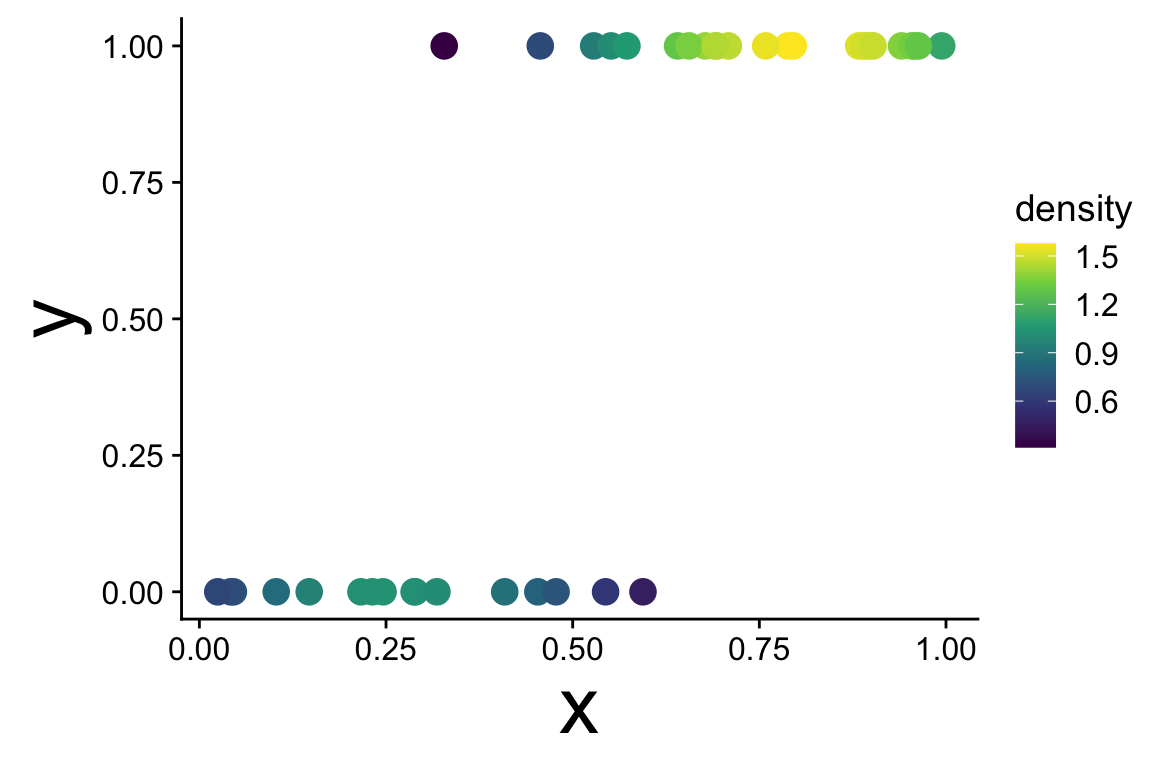
Recognizing a binomial distribution and a model bounded between [0, 1] makes more sense
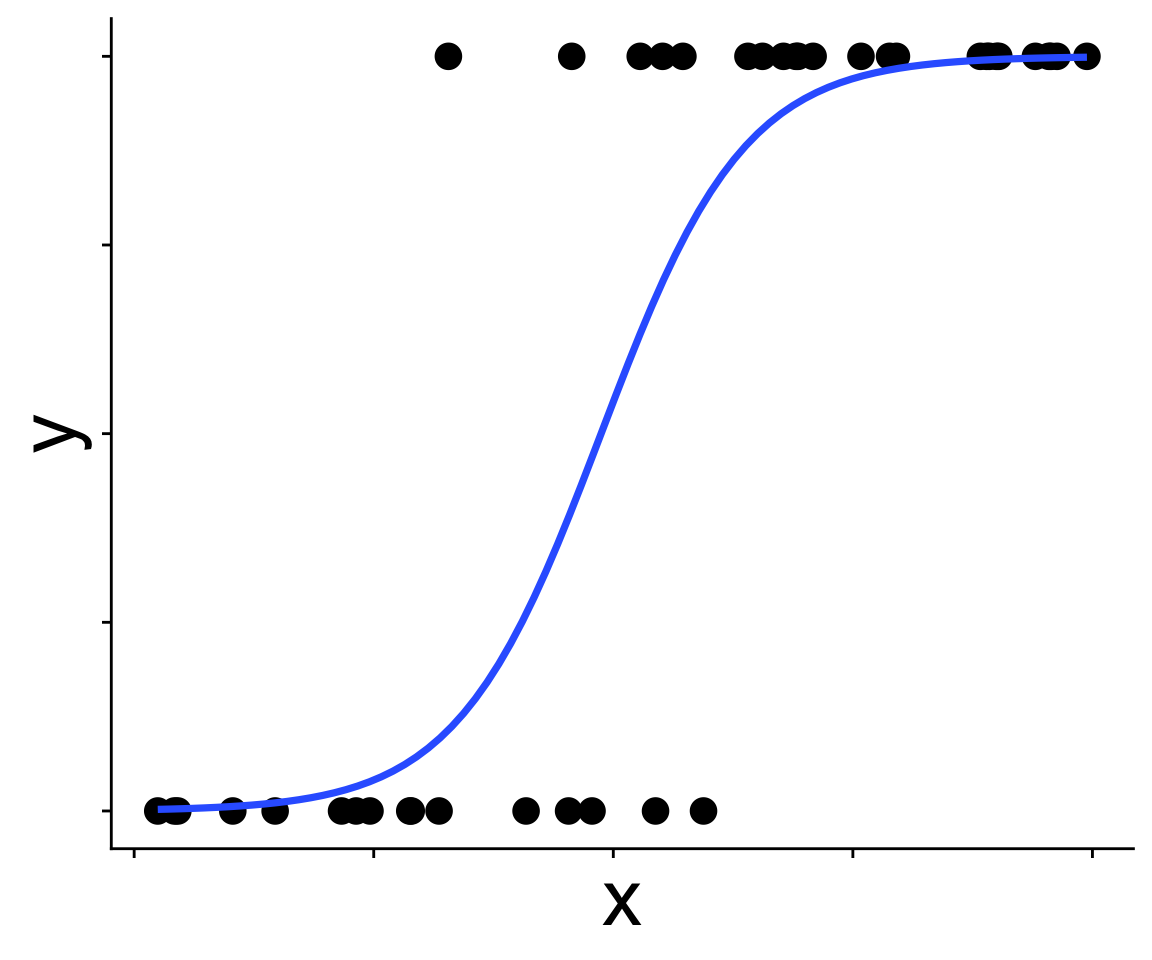
Binomial generalized linear models
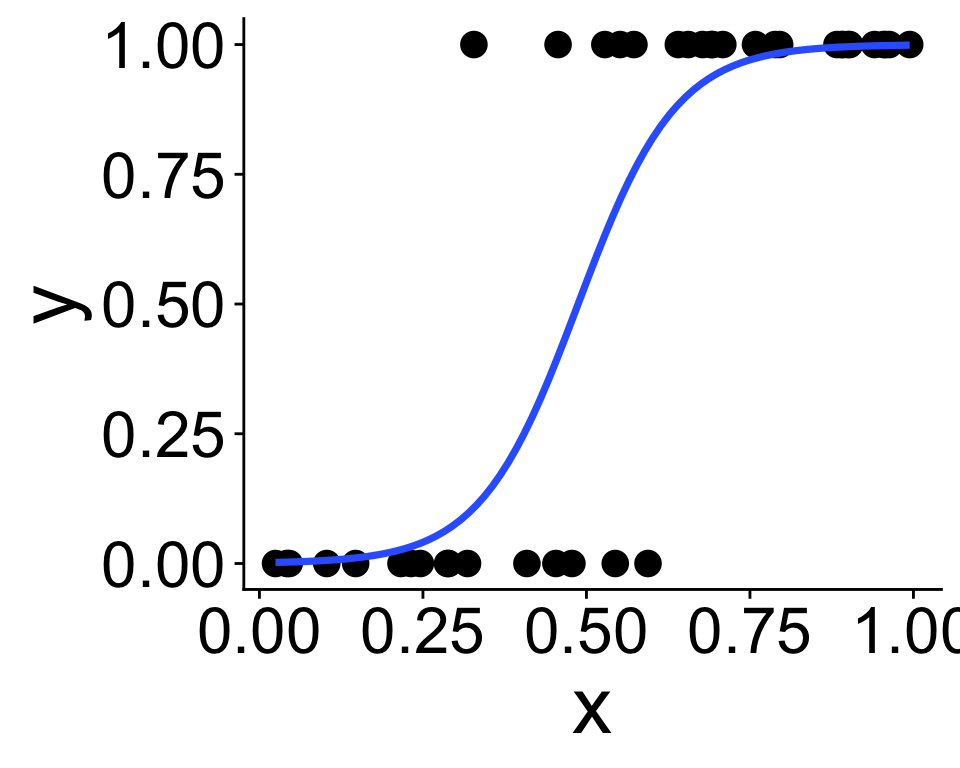
We’ve been here before: modeling presence/absence across a gradient using “expit” and “logit” functions
\[ p = \frac{1}{1 + e^{-(b_0 + b_1 x)}} \]
In GLM this kind of equation is called the link function
And we can use \(p\) in a binomial distribution
\[ y \sim \text{Binom}(p,~N = 1) \]
Understanding the link function
This is called the expit function
\[ p = \frac{1}{1 + e^{-(b_0 + b_1 x)}} \]
and produces the curve we just saw
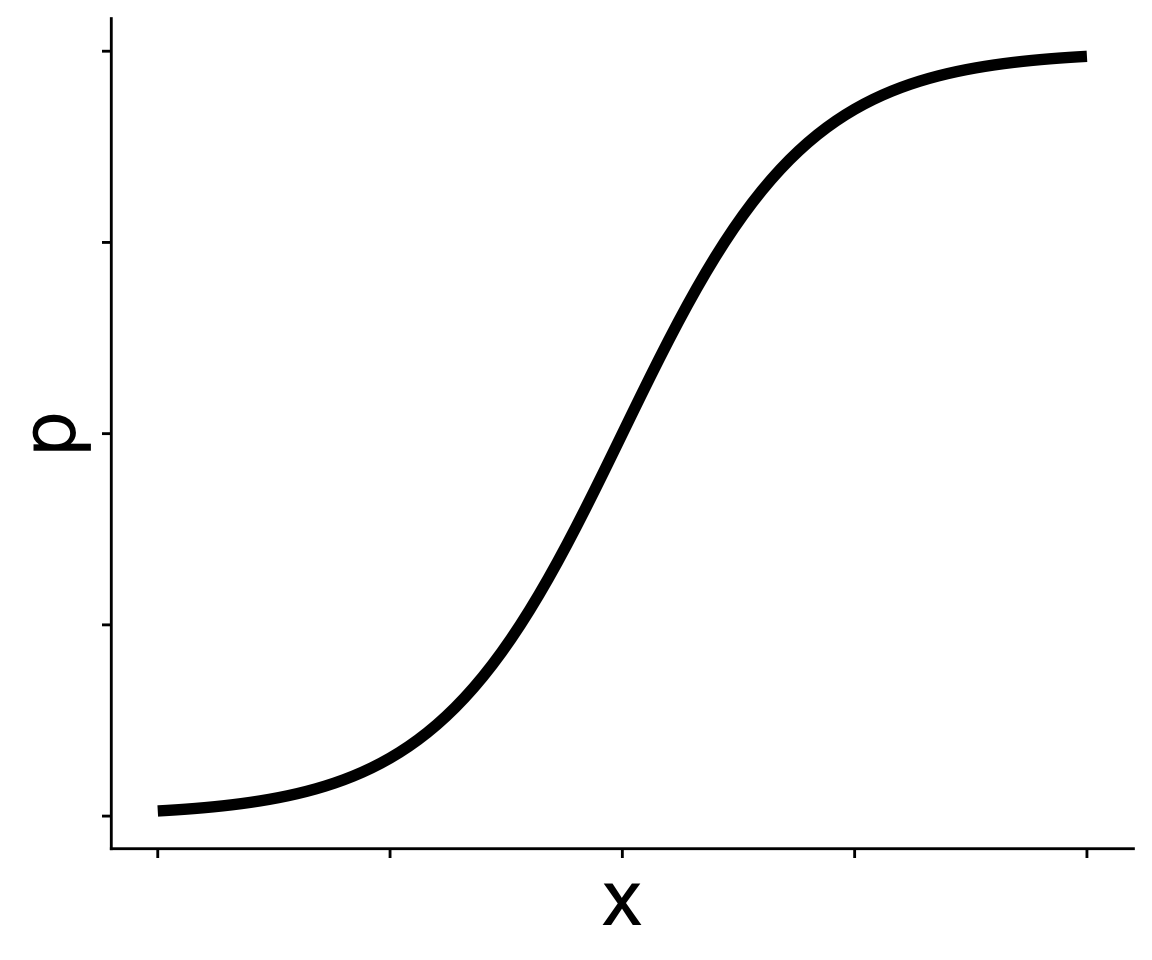
Understanding the link function
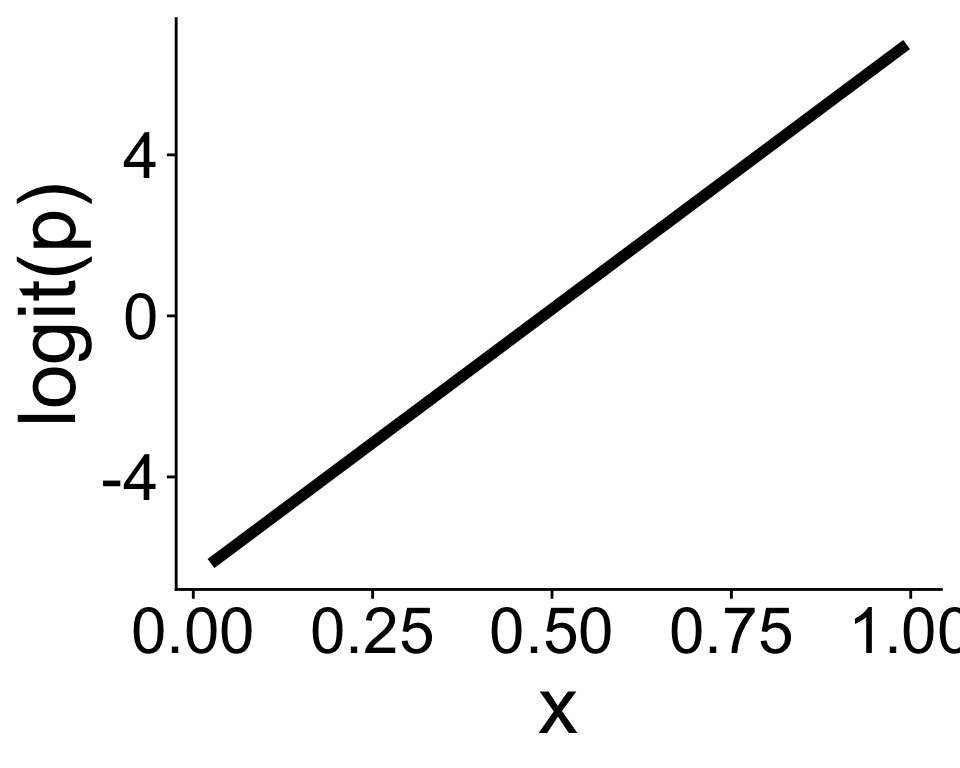
It’s often easier to think in terms of the logit function
\[ \text{logit}(p) = b_0 + b_1 x \]
which is the inverse of the expit function
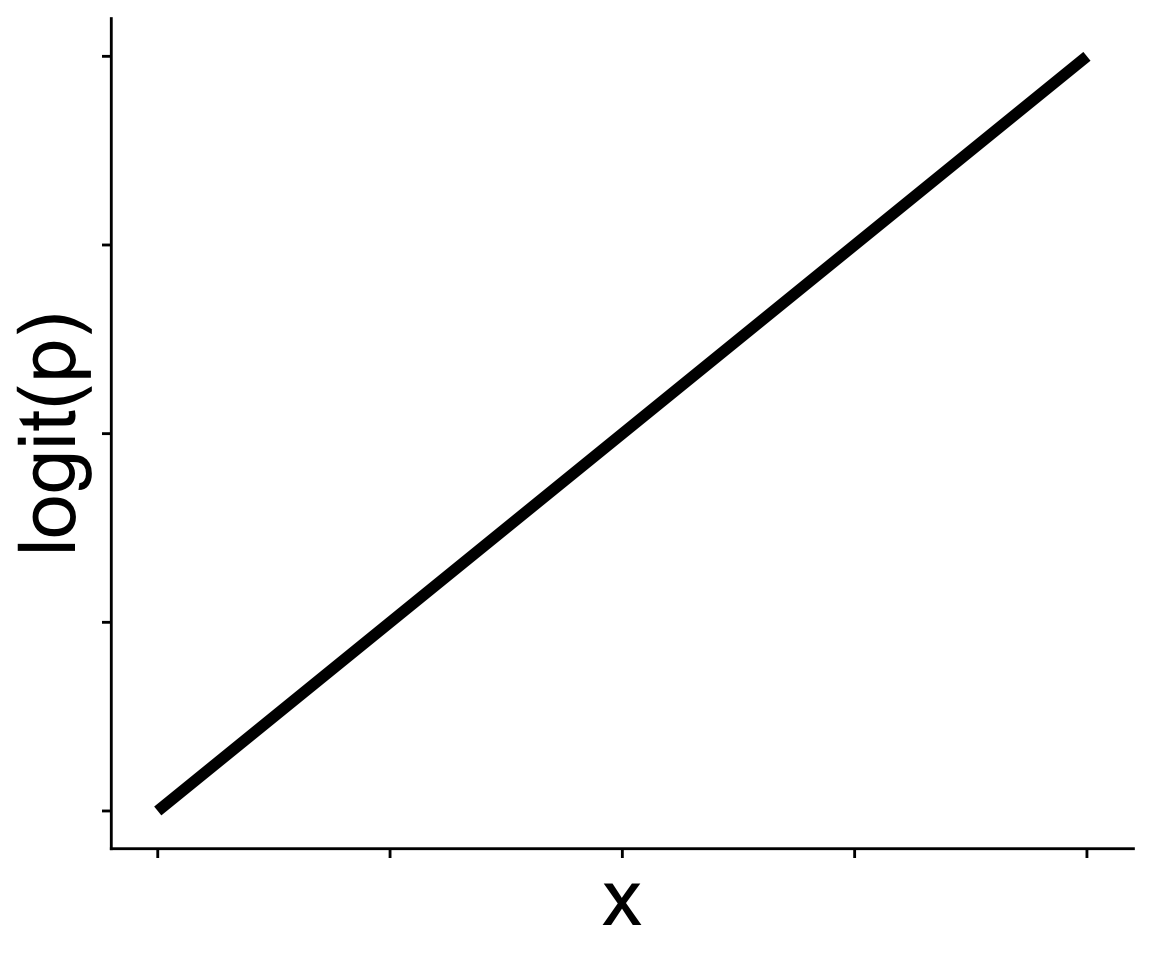
Now we just have a straight line! The link function gives us the linear in a generalized linear model
Generalized linear models in R: glm
How do we fit GLMs in R, why with the glm function!
Compared to lm there is one new thing: family = binomial
The argument family = binomial tells glm two things:
- we’re assuming a binomial distribution
- we want to use a logit link function (though other link functions are available)
Interpreting coefficients
Interpreting coefficients
Visualizing the model and data
ggplot2 is ready to help
(with some extra modification)
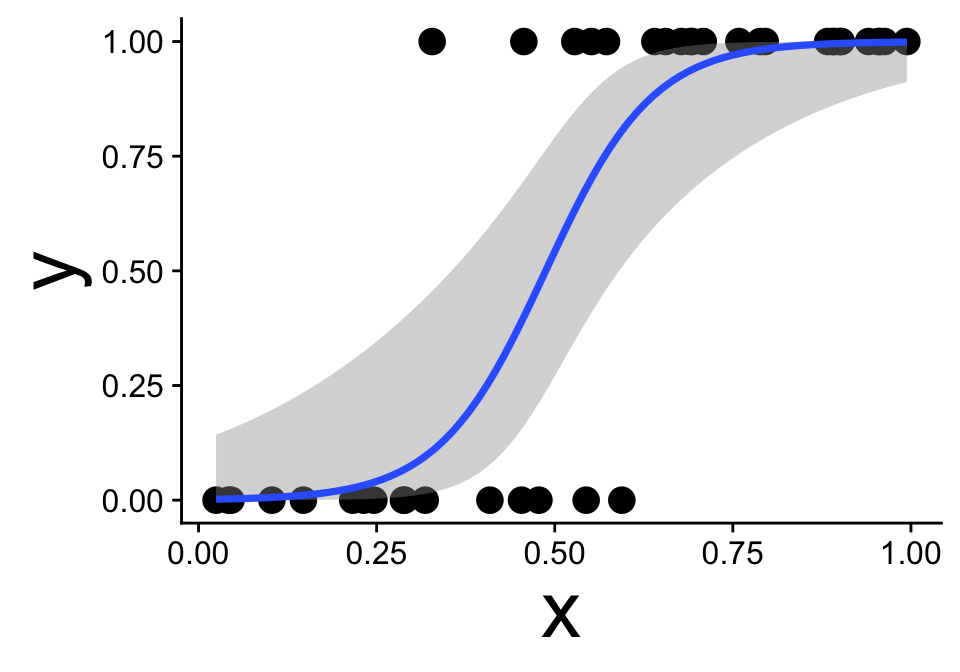
Visualizing the model and data
Visualizing the model and data
Visualizing the model and data
Visualizing the model and data
Let’s look at the link function in two ways
- With the expit equation
# get link function intercept and slope
bb <- coefficients(mod)
bb
# write a function for the equation
my_probit <- function(x, intercept, slope) {
intercept + slope * x
}
# use `geom_function`
ggplot(data.frame(x = 0:1, y = 0:1), aes(x, y)) +
geom_function(fun = my_probit,
args = list(intercept = bb[1],
slope = bb[2])) +
ylab("logit(p)")Visualizing the model and data
Let’s look at the link function in two ways
- With the predicted values for logit(p)
# function `predict` gives us model predictions
# `type = link` says, give us predictions on link function scale
logitp_hat <- predict(mod, type = "link")
head(logitp_hat)
# add a column for logit(p) and you're ready to plot!
ggplot(mutate(dat, logitp = logitp_hat),
aes(x, logitp)) +
geom_line(linewidth = 2) +
ylab("logit(p)")Visualizing the model and data
Either way gives you this graph
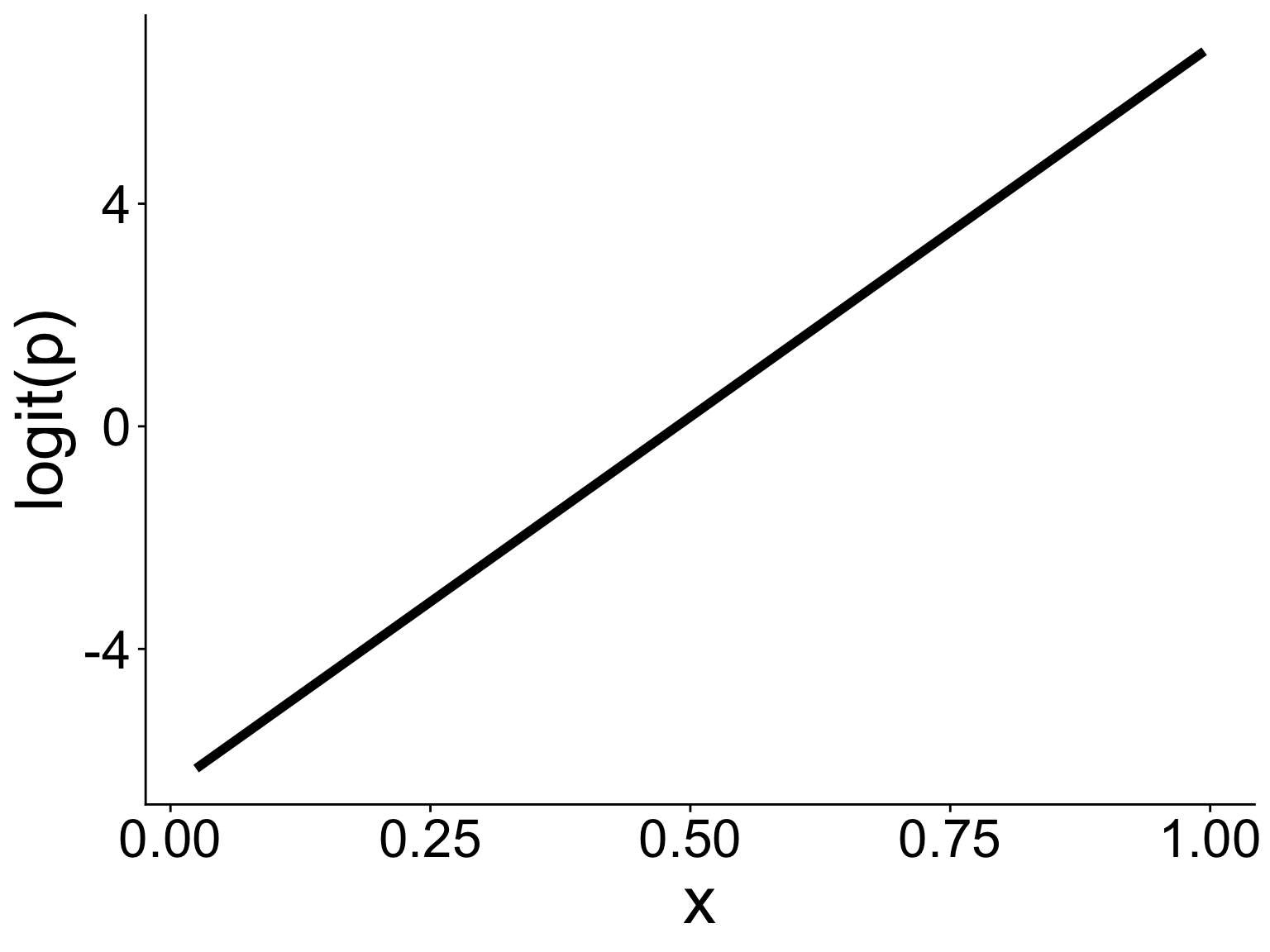
Binomial GLM: Use cases
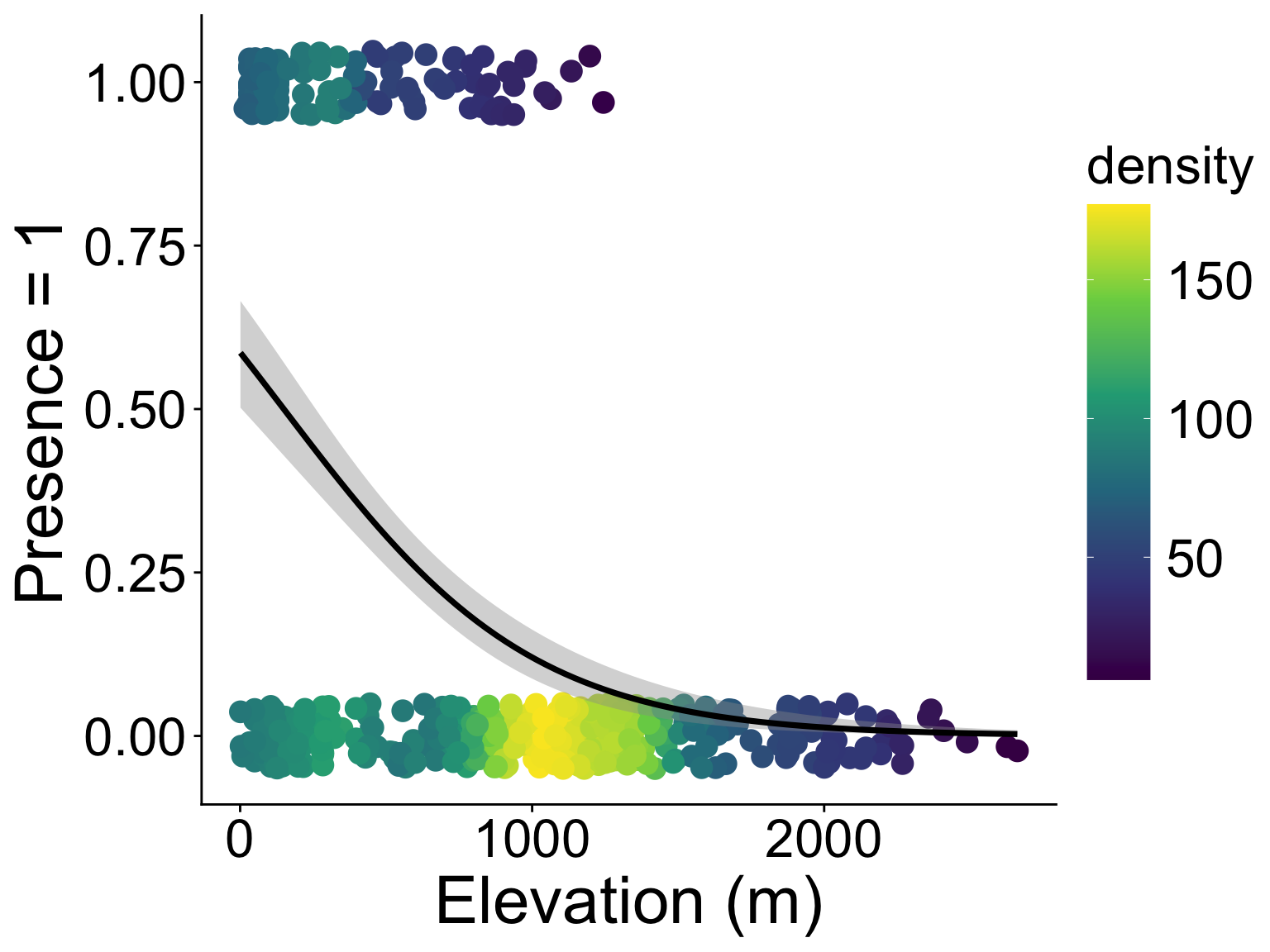
presence/absence, e.g., where is the strawberry guava?
Binomial GLM: Use cases
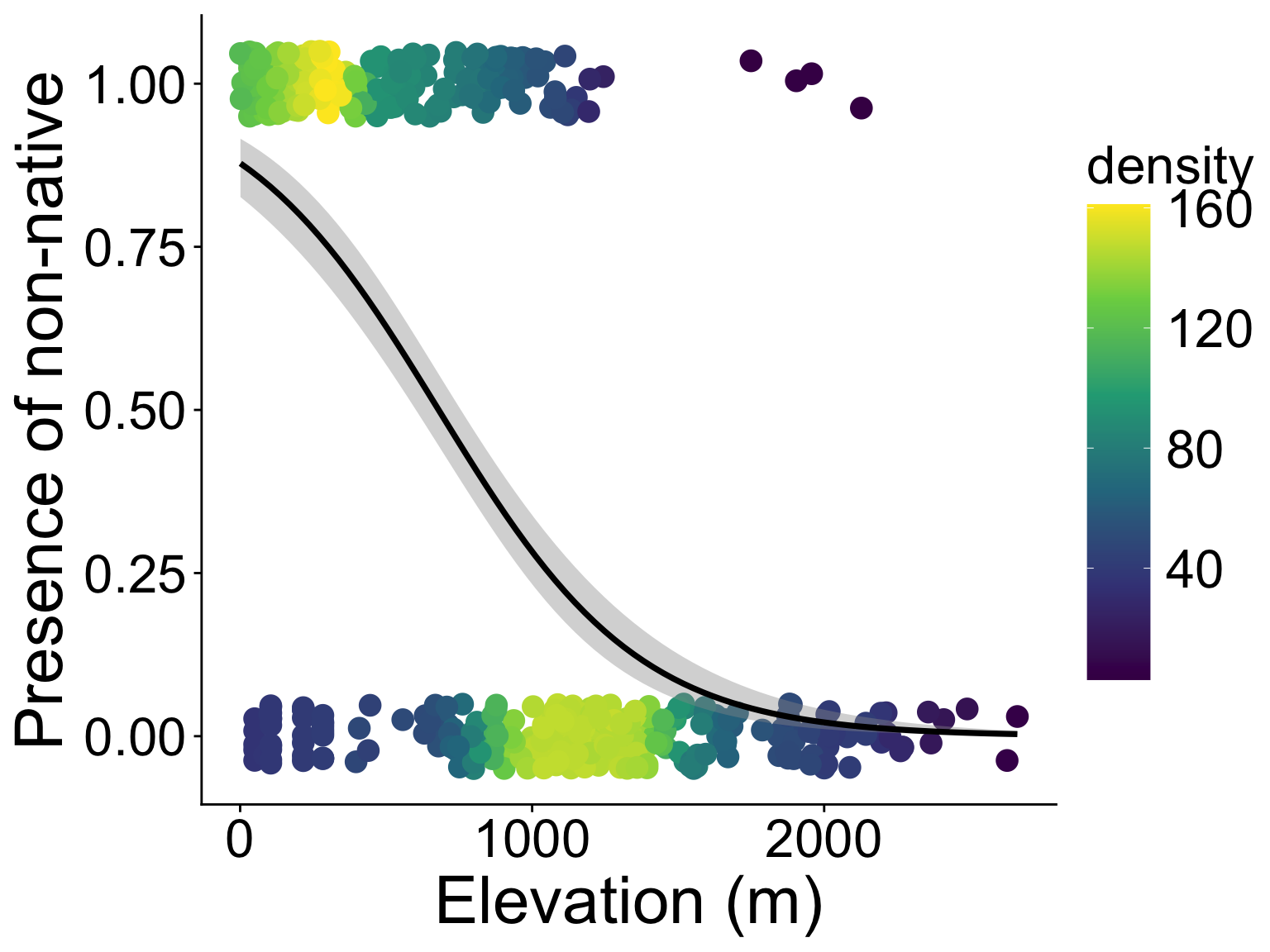
binary property, e.g., non-native or native?
Binomial GLM: Use cases
binary property, wait, aren’t we wasting tree-level data?
Binomial GLM: Use cases
binary property, e.g., non-native or native?
No that wasn’t a good idea—pseudo replication in plots
But before you go thinking of random effects
Remember, within each plot is like a binomial sample:
- “successes”: number of non-native trees
- total size: number of all trees
Binomial GLM: Use cases
number of successes vs. failures, back to plot-level data
| PlotID | elev_m | any_non | num_non | num_tre |
|---|---|---|---|---|
| 1 | 395.1798 | 1 | 9 | 75 |
| 2 | 129.6578 | 1 | 7 | 80 |
| 3 | 129.6578 | 1 | 1 | 37 |
| 4 | 129.6578 | 1 | 56 | 100 |
| 5 | 129.6578 | 1 | 49 | 99 |
| 6 | 129.6578 | 1 | 23 | 64 |
Binomial GLM: Use cases
number of successes vs. failures, back to plot-level data
ggplot(tree_plot, aes(elev_m, num_non / num_tre,
# include `success` and `failure`
# for use in `geom_smooth`
success = num_non,
failure = num_tre - num_non)) +
geom_pointdensity() +
scale_color_viridis_c() +
geom_smooth(method = "glm", color = "black",
# need to add a specific formula
formula = cbind(success, failure) ~ x,
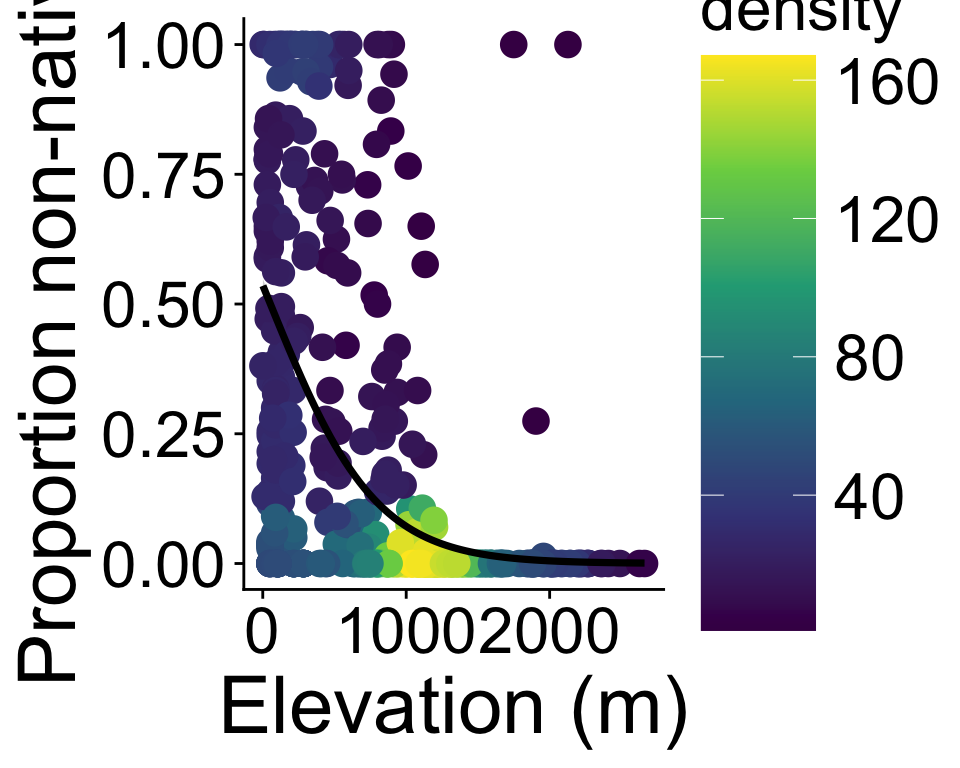
method.args = list(family = binomial))Binomial GLM: Use cases
number of successes vs. failures, back to plot-level data
(with some extra beautifying)