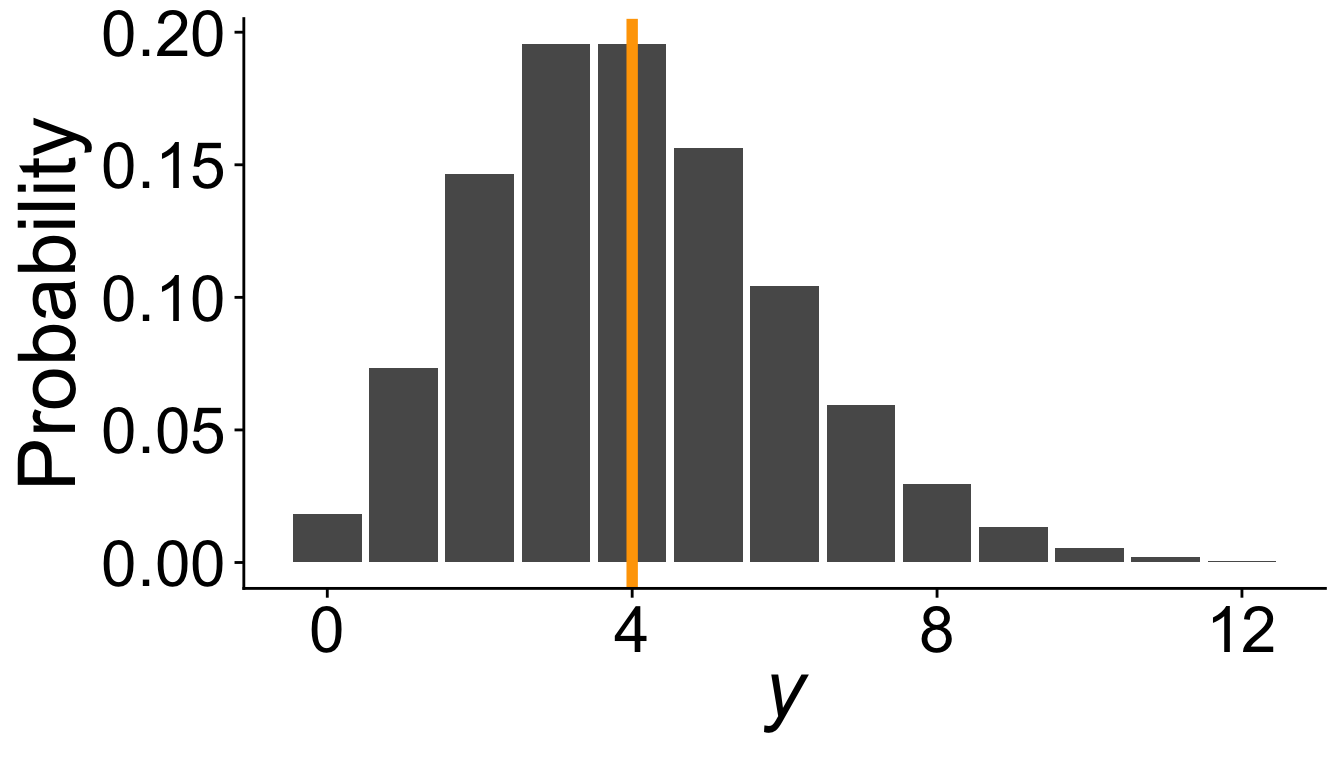
The Poisson Generalized Linear Model
Consider the Poisson distribution…
\[ y \sim \text{Pois}(\lambda) \]
Often used to model counts of things (number of trees, number of species, etc.)
\(\lambda\) is both the mean and the variance of the distribution
Consider the Poisson distribution…
\(\lambda\) is both the mean and the variance of the distribution.
What must be true about \(\mathbf{\lambda}\)?
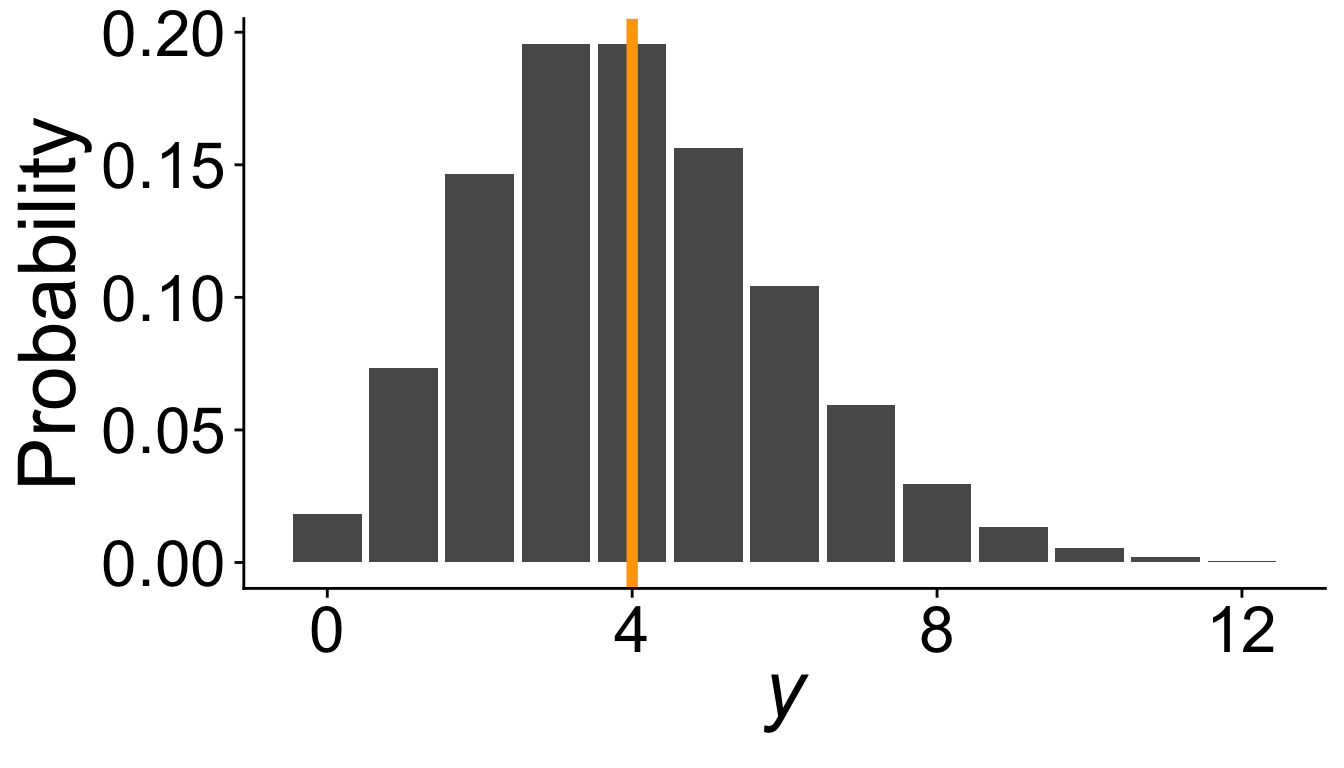
must be true: \(\lambda \geq 0\)
Consider the Poisson GLM…
If we wanted to use a Poisson distribution in a GLM, we have to make sure \(\lambda \geq 0\)
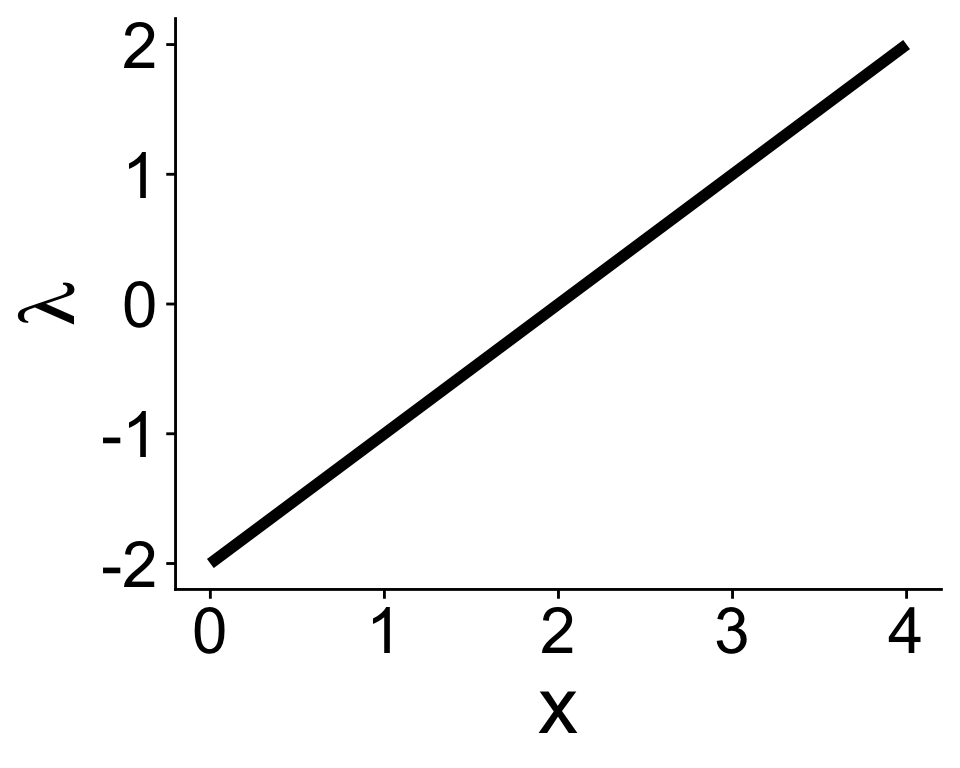
Can’t have this
\[ \lambda = b_0 + b_1 x \]
Need some kind of link function
Consider the Poisson GLM…
If we wanted to use a Poisson distribution in a GLM, we have to make sure \(\lambda \geq 0\)
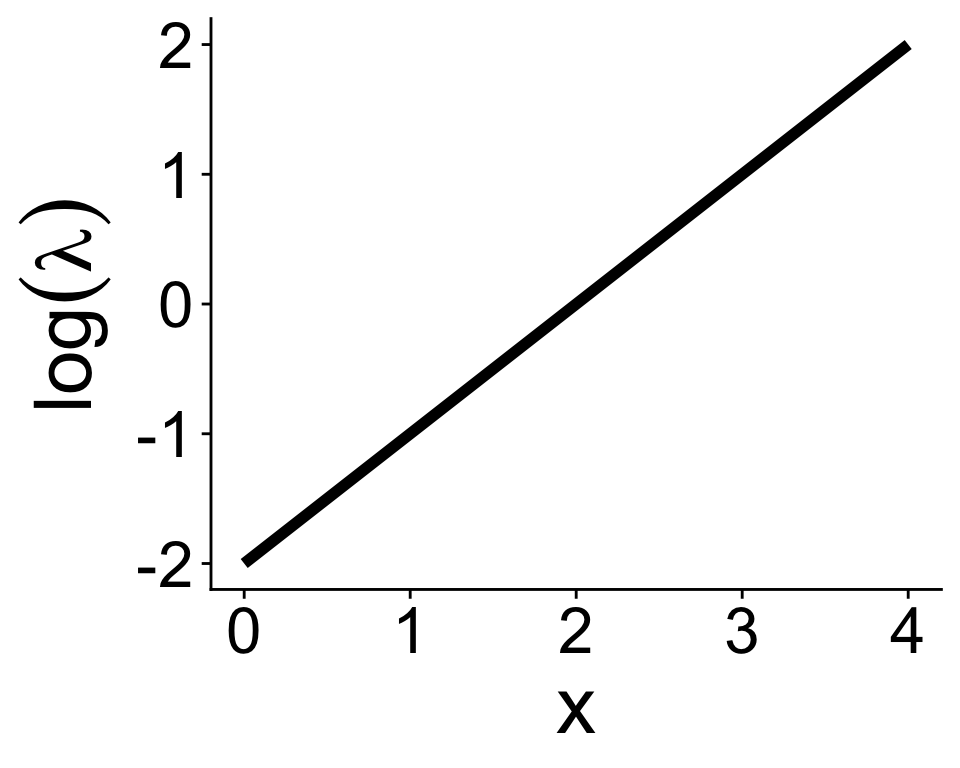
\(e^\text{anything} \geq 0\), always
\[ \lambda = e^{b_0 + b_1 x} \]
And the inverse gives us back the straight line
\[ \log(\lambda) = b_0 + b_1 x \]
So with Poisson GLM, a log link function is the default
Poisson GLM with glm function in R
Back to species area
\[ \begin{aligned} S &= cA^z \\ \log(S) &= \log(c) + z\log(A) \end{aligned} \]
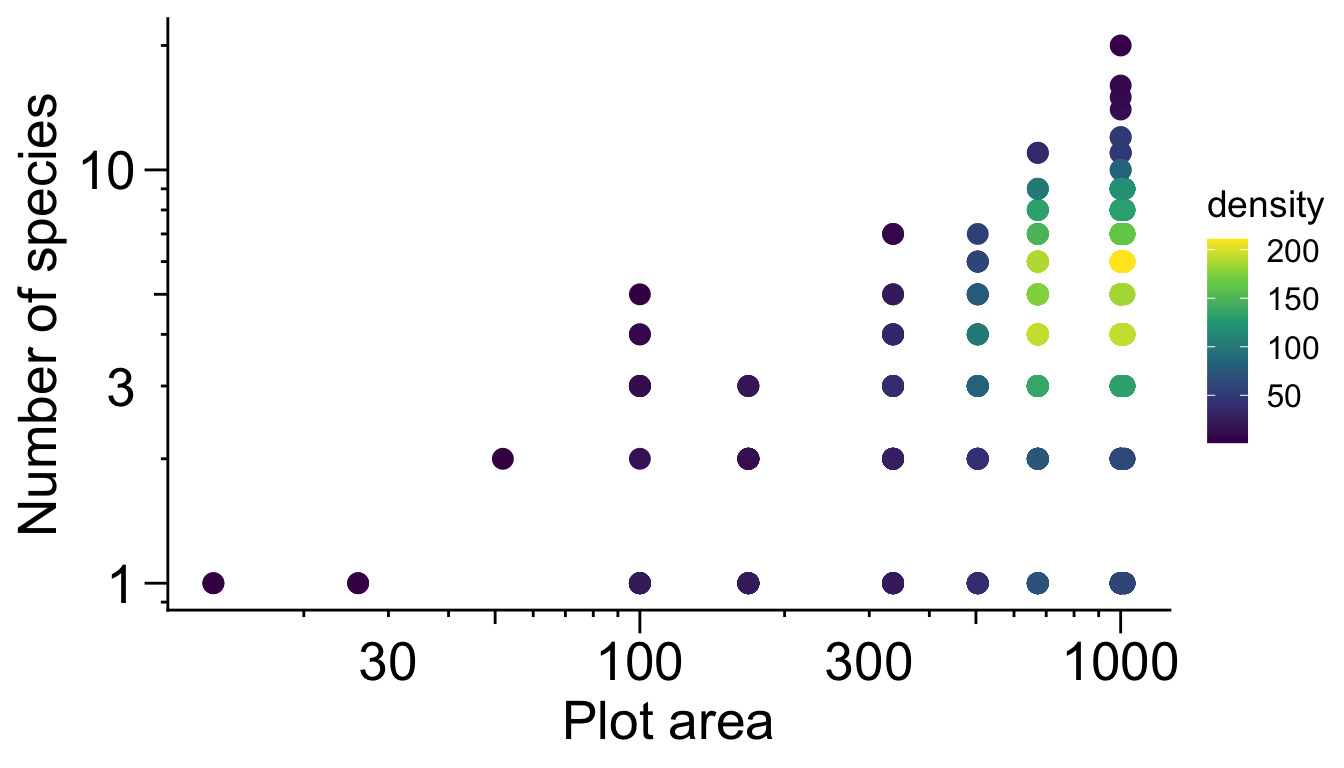
(note: 2 super large plots excluded)
lends itself to a log link function
\[ \begin{aligned} \bar{S} &= \lambda \\ \log(A) &= x \\ \Rightarrow \log(\bar{S}) &= b_0 + b_1 \log(A) \end{aligned} \]
Poisson GLM with glm function in R
with data like these
| Plot_Area | nspp |
|---|---|
| 1000.00 | 9 |
| 1017.88 | 9 |
| 1017.88 | 2 |
| 1017.88 | 9 |
| 1017.88 | 4 |
| 1017.88 | 9 |
What besides area?
- We are probably interested in more than area
- Area might even be a “nuisance variable”
- one that we have to account for
- but we’re more interested in other variables
- How about
hii(human impact) andavg_temp_annual_c(temperature)? - Can!
Adding more explanatory variables
(Intercept) log(Plot_Area) hii avg_temp_annual_c
-2.320992411 0.516567381 0.011011301 0.007295895 After accounting for area, species richness increases (weakly) with both human impact and temperature
Perhaps non-native plants augment species richness (Sax & Gaines 2003)
Or perhaps lowland (i.e. hot) forests were incredibly diverse (Rock 1913) and are now lost to heavy human modification
Adding more explanatory variables
Beyond biological interpretation, adding more variables requires care
This is the likelihood surface for our model, visualizing just the coefficients for human impact and temperature
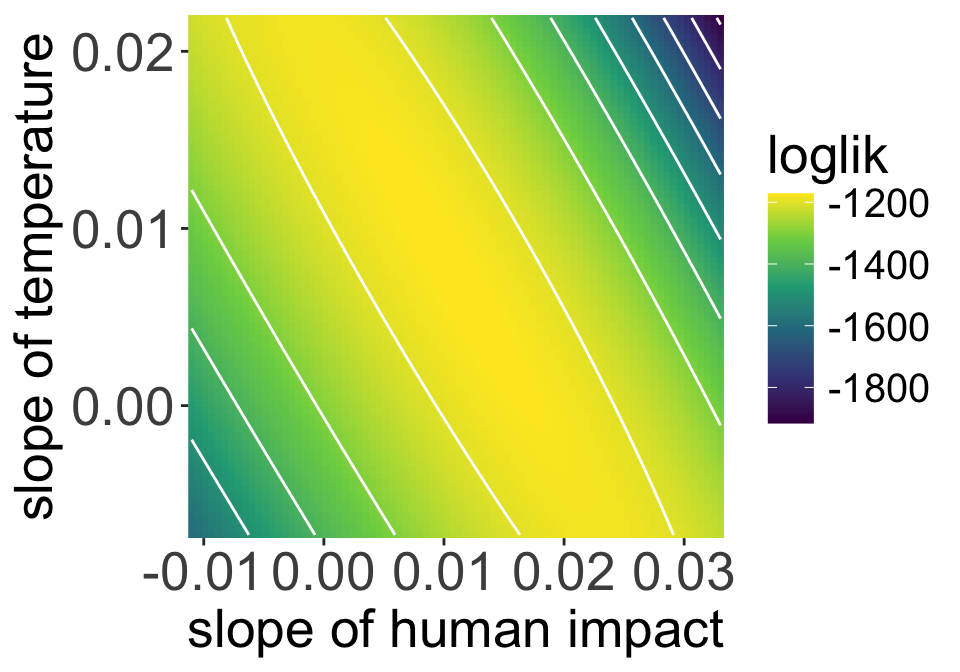
There is a long, flat ridge across the likelihood surface
That’s a problem because it is difficult and unreliable to figure out the optimal parameter combination
why?
Adding more explanatory variables
Collinearity creates these kinds of ridges
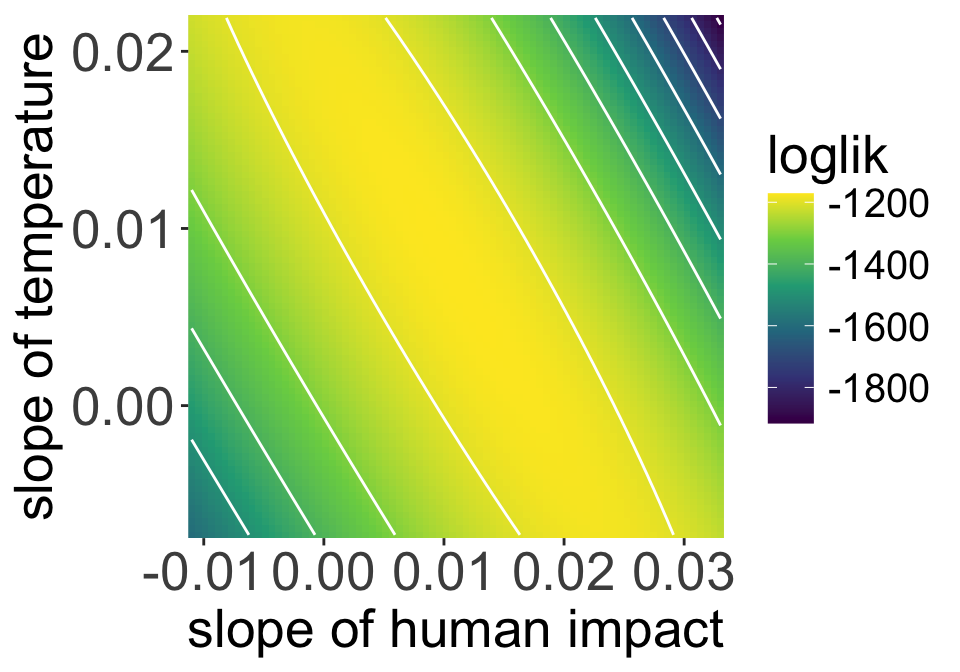
Human impact and temperature are correlated
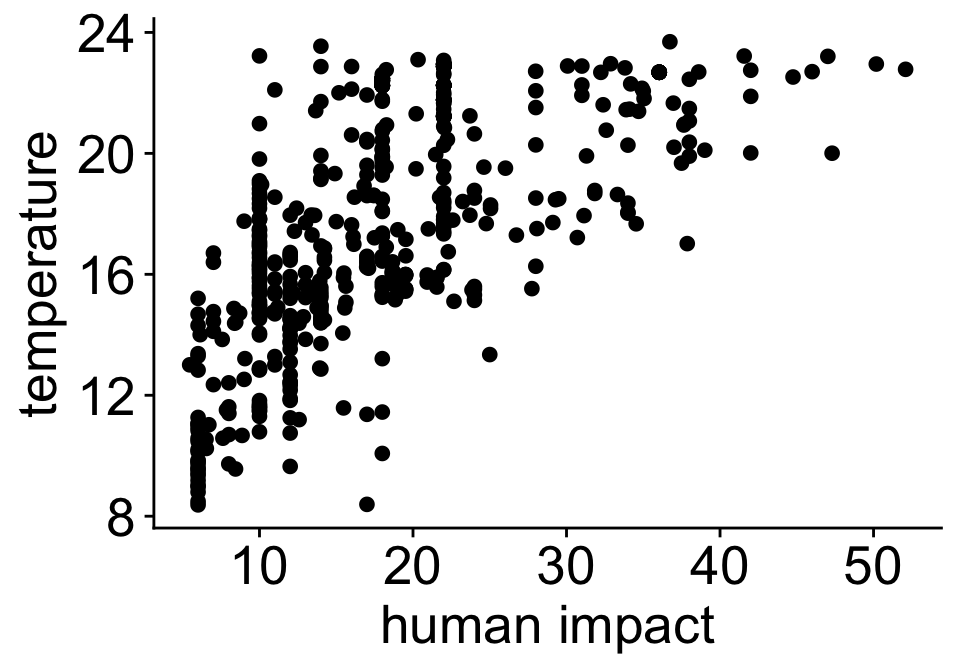
Check for collinearity, a common cut off is \(-0.6 < r < 0.6\)